Sharper, Deeper, Faster: Interdisciplinary Team Develops Advanced Live-Imaging Approach
(Originally published by Caltech)
July 25, 2011
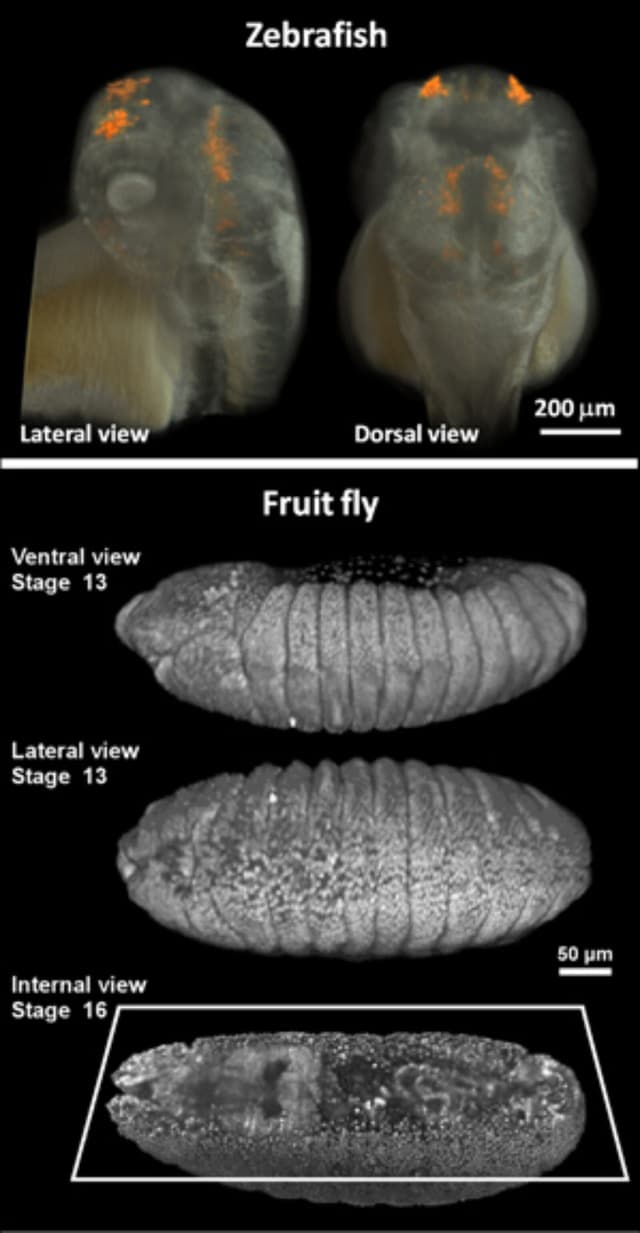
![Three-dimensional live imaging of zebrafish (upper panel) and fruit fly (lower panel) embryos with two-photon light sheet microscopy. The upper panel shows not only the embryo's tissue (in white) but also green fluorescent protein driven by a gene-specific promoter (in orange). The lower panel shows the individual cell nuclei of the fruit fly embryo, labeled with green fluorescent protein, imaged at different time points during its embryonic development.[Credit: Willy Supatto, Seth Ruffins, Thai Truong / Caltech]](/uploads/images/image/nanoscience/1823-Small-Fraser_Truong_Image-Zebrafish_Fly_medium.jpg)
For modern biologists, the ability to capture high-quality, three-dimensional (3D) images of living tissues or organisms over time is necessary to answer problems in areas ranging from genomics to neurobiology and developmental biology. The better the image, the more detailed the information that can be drawn from it. Looking to improve upon current methods of imaging, researchers from the California Institute of Technology (Caltech) have developed a novel approach that could redefine optical imaging of live biological samples by simultaneously achieving high resolution, high penetration depth (for seeing deep inside 3D samples), and high imaging speed.
The imaging technique is explained in a paper in the advance online publication of the journal Nature Methods, released on July 14. It will also appear in an upcoming print version of the journal.
"Before our work, the state-of-the-art imaging techniques typically excelled in only one of three key parameters: resolution, depth, or speed. With our technique, it's possible to do well in all three and, critically, without killing, damaging, or adversely affecting the live biological samples," says biologist Scott Fraser, director of the Biological Imaging Center at Caltech's Beckman Institute, board member of Caltech's Kavli Nanoscience Institute, and senior author of the study.
The research team achieved this imaging hat trick by first employing an unconventional imaging method called light-sheet microscopy, where a thin, flat sheet of light is used to illuminate a biological sample from the side, creating a single illuminated optical section through the sample. The light given off by the sample is then captured with a camera oriented perpendicularly to the light sheet, harvesting data from the entire illuminated plane at once. This allows millions of image pixels to be captured simultaneously, reducing the light intensity that needs to be used for each pixel. This not only enables fast imaging speed but also decreases the light-induced damage to the living samples, which the teams demonstrated using the embryos of fruit fly and zebrafish.
To achieve sharper image resolution with light-sheet microscopy deep inside the biological samples, the team used a process called two-photon excitation for the illumination. This process has been used previously to allow deeper imaging of biological samples; however, it usually is used to collect the image one pixel at a time by focusing the exciting light to a single small spot.
"The conceptual leap for us was to realize that two-photon excitation could also be carried out in sheet-illumination mode," says Thai Truong, a postdoctoral fellow in Fraser's laboratory and first author of the paper. This novel side-illumination with a two-photon illumination is the topic of a pending patent.
"With this approach, we believe that we can make a contribution to advancing biological imaging in a meaningful way," continues Truong, who did his PhD training in physics. "We did not want to develop a fanciful optical imaging technique that excels only in one niche area, or that places constraints on the sample so severe that the applications will be limited. With a balanced high performance in resolution, depth, and speed, all achieved without photo-damage, two-photon light-sheet microscopy should be applicable to a wide variety of in vivo imaging applications." He credits this emphasis on wide applicability to the interdisciplinary nature of the team—which includes two biologists, two physicists, and one electrical engineer.
"We believe the performance of this imaging technique will open up many applications in life sciences and biomedical research—wherever it is useful to observe, non-invasively, dynamic biological process in 3D and with cellular or subcellular resolution," says Willy Supatto, co–author of the paper and a former postdoctoral fellow in Fraser's laboratory (now at the Centre National de la Recherche Scientifique, in France).
One example of such an application would be to construct 3D movies of the entire embryonic development of an organism, covering the entire embryo in space and time. These movies could capture what individual cells are doing, as well as important genes' spatiotemporal expression patterns—elucidating the activation of those genes within specific tissues at specific times during development.
"The goal is to create 'digital embryos,' providing insights into how embryos are built, which is critical not only for basic understanding of how biology works but also for future medical applications such as robotic surgery, tissue engineering, or stem-cell therapy," says Fraser. The team's first attempt at this can be seen in the accompanying movie, in which the cell divisions and movements that built the entire fruit fly embryo were captured without perturbing its development.
The Nature Methods paper is titled "Deep and fast live imaging with two-photon scanned light-sheet microscopy." David Koos, senior research scientist at Caltech's Beckman Institute, and John Choi, a former postdoctoral fellow in Fraser's laboratory, also contributed to the study.
The research was supported by the Beckman Institute and the U.S. National Institutes of Health.